Author Affiliations
Abstract
Background: Cancer remains a leading global health challenge, with nearly 10 million deaths in 2020. Traditional therapies like chemotherapy and radiation often face limitations, including systemic toxicity, acquired resistance, and suboptimal tumor targeting. The advent of Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-Cas9 technology has revolutionized genetic research by enabling precise gene editing. Its adaptation for epigenetic modulation presents novel opportunities to reversibly regulate gene expression in cancer without introducing permanent deoxyribonucleic acid (DNA) alterations.
Objective: This literature review evaluates the therapeutic potential of CRISPR-based epigenetic editing in cancer treatment and examines recent clinical studies that inform its translational prospects.
Methodology: A systematic literature search was conducted using PubMed for clinical trials and randomized controlled trials published between January 2020 and June 2025, focusing on CRISPR, epigenetics, and cancer therapy. Five clinical studies met the inclusion criteria and were reviewed following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) 2020 guidelines.
Results: The reviewed studies highlight genetic mutations and epigenetic alterations in cancers such as human epidermal growth factor receptor 2 (HER2)-mutant breast cancer, prostate cancer, and peripheral T-cell lymphomas. CRISPR-dCas9 fusion tools and ribonucleic acid (RNA)-targeting systems like Cas13 offer precision in modulating gene expression, potentially overcoming drug resistance and reactivating tumor suppressors. However, efficient delivery, specificity, and safety remain significant barriers.
Conclusion: CRISPR-mediated epigenetic editing holds transformative potential in precision oncology by enabling targeted, reversible gene regulation. Future research must address delivery technologies, off-target risks, and clinical validation to advance CRISPR applications in cancer therapeutics.
Keywords
Clustered regularly interspaced short palindromic repeats, Epigenetic editing, Cancer therapy, Gene regulation, Precision oncology.
Introduction
Cancer remains one of the leading causes of mortality worldwide, accounting for nearly 10 million deaths in 2020, with an estimated 2 to 2.5 million cases prevalent in India. The complexity of cancer treatment, often involving multimodal approaches such as chemotherapy, radiation, and surgery, exacerbates the risk of drug-related problems, contributing to therapeutic complications and reduced patient quality of life.[1]
In the pursuit of more targeted and effective treatments, the advent of CRISPR-Cas9 genome editing has revolutionized biomedical research. Recognized for its high precision, versatility, and simplicity, CRISPR-Cas9 enables targeted genetic modifications that have immense therapeutic potential, particularly in cancer treatment. The foundational mechanism of CRISPR-Cas9 relies on a single-guide RNA (sgRNA) that directs the Cas9 endonuclease to specific genomic loci, inducing double-stranded DNA breaks (DSBs). These DSBs are subsequently repaired via endogenous cellular mechanisms, including non-homologous end joining (NHEJ) or homology-directed repair (HDR), facilitating precise gene editing.[2-5]
Compared to previous genome editing platforms such as zinc-finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs), CRISPR-Cas9 offers superior accuracy and flexibility, largely due to the straightforward Watson-Crick base pairing between the sgRNA and the target DNA sequence.[6,7] The simplicity of CRISPR’s design allows for multiplexed gene editing, enabling simultaneous targeting of multiple genes. Moreover, CRISPR-based technologies have evolved to modulate gene expression without permanently altering the genome. Techniques such as CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) employ catalytically dead Cas9 (dCas9) fused with transcriptional activators or repressors to upregulate or downregulate gene expression, respectively.[8-10]
Beyond DNA editing, CRISPR tools have expanded to RNA targeting, with engineered Cas9 variants enabling post-transcriptional gene silencing, RNA imaging, and interventions against RNA viruses. These capabilities extend the therapeutic horizon of CRISPR beyond permanent DNA alterations, providing transient and reversible regulatory mechanisms suitable for cancer treatment.[11-15] For instance, RNA-targeting Cas systems like Cas13 have demonstrated efficacy in modulating gene expression at the transcriptomic level, which is particularly relevant for cancers driven by aberrant RNA splicing or non-coding RNAs.
However, the clinical translation of CRISPR technologies is not without challenges. Efficient and targeted delivery of CRISPR components to specific cells or tissues in vivo remains a formidable hurdle. Current delivery systems, including viral vectors (like AAV and lentivirus) and non-viral methods (lipid nanoparticles), each present limitations in terms of immunogenicity, cargo capacity, and cell-type specificity.[16,17] Furthermore, the risk of off-target effects, unintended modifications at non-specific genomic sites, poses significant safety concerns that must be mitigated before widespread therapeutic application can be realized.[18,19]
Conventional cancer therapies, while having improved survival outcomes in some malignancies, are often associated with systemic toxicity, acquired resistance, and suboptimal targeting of tumor cells.[20] For example, despite the efficacy of temozolomide in glioblastoma, the five-year survival remains dismal, underscoring the need for more precise interventions.[21] High-throughput sequencing and multi-omics analyses have revealed the intricate genetic and epigenetic landscape of cancers, highlighting mutations, gene duplications, and RNA alterations that contribute to tumorigenesis and progression.[22] With over 140 cancer-associated gene mutations characterized, tailoring effective therapies remains complex and necessitates innovative approaches.[23]
Epigenetic alterations, heritable yet reversible changes in gene expression that do not involve alterations in the DNA sequence, play a critical role in cancer initiation and progression. Epigenetic mechanisms include DNA methylation, histone modifications, chromatin remodeling, and the regulation by non-coding RNAs like microRNAs (miRNAs). These mechanisms can silence tumor suppressor genes or activate oncogenes, driving malignant transformation.[24] The human genome, comprising approximately 20,000 genes across 23 chromosome pairs, generates a diverse transcriptome of coding and non-coding RNAs, each playing distinct roles in cellular homeostasis and disease states, including cancer.[25]
The completion of the Human Genome Project catalyzed the development of genome editing technologies, including RNA interference (RNAi), ZFNs, and TALENs. While these tools enabled targeted gene disruption, their utility in addressing complex epigenetic modifications was limited by issues of efficiency, specificity, and off-target effects.[26] CRISPR-Cas9, derived from bacterial adaptive immunity against invading phages, emerged as a more versatile and robust tool capable of not only correcting genetic mutations but also modulating the epigenome.[10]
CRISPR’s integration with epigenetic regulators offers a precise approach to cancer therapy by modifying gene expression without causing DNA breaks, enabling the reactivation of tumor suppressors or silencing oncogenes with reduced genomic risk. dCas9-based tools can also reprogram malignant cells to less aggressive states and help model cancer epigenetics for research. Despite advancements, optimizing guide RNA specificity and minimizing off-target effects remain challenges, but artificial intelligence and deep learning are enhancing sgRNA prediction and precision. Emerging CRISPR systems like Cas12 and Cas13 further expand DNA and RNA targeting capabilities, supporting therapies for non-coding RNA-driven cancers. Collectively, these innovations position CRISPR as a transformative tool in precision oncology, though hurdles in delivery, specificity, and ethics must still be addressed.
The aim of this literature review is to synthesize recent clinical evidence on the application of CRISPR-based epigenetic modulation in cancer therapy, identify therapeutic opportunities, and outline current limitations and future research directions in this rapidly evolving field.
Methodology
A systematic literature search was performed using PubMed databases to identify relevant clinical trials and randomized controlled trials (RCTs) investigating the application of CRISPR-based gene editing in conjunction with epigenetic modulation for cancer therapy. The search strategy employed the terms (CRISPR OR “CRISPR-Cas9”) AND (Epigenetics OR “Histone Deacetylase” OR HDAC OR Entinostat) AND (Cancer OR Tumor OR Neoplasm) AND (“Gene Editing” OR Therapy)
Search filters were applied for:
- Publication period: January 1, 2020, to June 30, 2025
- Study type: Clinical trials and randomized controlled trials
- Subjects: Human participants (both male and female)
- Language: English
Inclusion Criteria
- Clinical trials or randomized controlled trials.
- Human participant studies
- Studies published in English within the last 5 years.
- Studies including both male and female participants.
Exclusion Criteria
- Editorials, commentaries, letters, book chapters, and conference abstracts without peer-reviewed data.
- Case reports, case series, and narrative or systematic reviews.
- In vitro or animal model studies.
- Non-English publications.
- Studies without available results sections.
- Articles not directly addressing CRISPR, gene editing, epigenetic therapies, or cancer treatment.
A systematic literature search was conducted to assess the effectiveness of Curcuma longa on inflammation. The search was performed across electronic databases, which yielded a total of 638 records. Before screening, 435 records were removed due to duplication, irrelevance, or other exclusion criteria applied at the initial stage. Following this, 203 records were screened based on their titles and abstracts. During this process, 120 records were excluded as they did not meet the predefined inclusion criteria related to the research objective.
The remaining 83 reports were sought for full-text retrieval. However, 55 reports could not be retrieved due to access limitations or unavailability of full-text versions. Consequently, 28 full-text articles were assessed for eligibility against detailed inclusion and exclusion criteria. A quality appraisal was performed using the Scale for the Assessment of Narrative Review Articles (SANRA) tool, where articles scoring less than 10 were excluded. As a result, 23 reports were excluded based on their SANRA score.
Ultimately, five studies met all eligibility criteria and were included in the final review. The entire study selection process adhered to the PRISMA 2020 guidelines. A visual summary of the selection process is provided in the PRISMA flow diagram, ensuring a transparent and reproducible methodology (Figure 1).
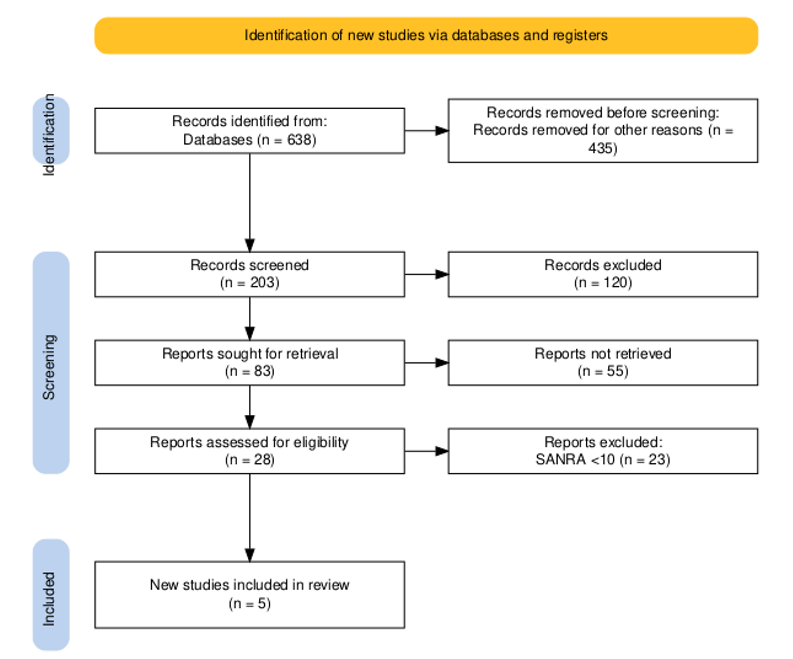
Figure 1: PRSIMA flow diagram
Results
In a Phase 2 clinical trial of Okines et al., 31 women with heavily pretreated HER2-mutated metastatic breast cancer (HER2-mut MBC) were enrolled between September 2021 and December 2022, with a median follow-up of 15 months. The median age was 64 years; 58% had lobular carcinoma, 42% ductal carcinoma, and 87% were hormone receptor positive. The study evaluated a chemotherapy-free combination of tucatinib and trastuzumab, achieving a confirmed objective response rate (cORR) of 41.9% (90% confidence interval [CI]: 26.9–58.2%), including two complete responses. Secondary endpoints demonstrated a median duration of response of 12.6 months, median progression-free survival (PFS) of 9.5 months (90% CI: 5.4–13.8), and median overall survival (OS) of 20.1 months. Disease control was observed in 80.6% of participants, with tumour shrinkage reported in 73%.
Exploratory biomarker analyses revealed HER2 mutations predominantly localized to the tyrosine kinase and extracellular domains, notably involving Leu755 and Ser310 mutations. These mutational hotspots represent promising targets for CRISPR-Cas9 gene editing or epigenetic modulation using CRISPR-dCas9 platforms to interrogate drug sensitivity and resistance mechanisms. Additionally, co-occurring mutations, such as CDH1 and PIK3CA, were identified; CDH1 alterations correlated with higher response rates (62.5%), whereas PIK3CA mutations were associated with poorer outcomes. These insights suggest potential roles for CRISPR-mediated gene editing to correct or silence resistance-associated mutations. Moreover, CRISPR-dCas9-based epigenetic editors, such as those fused with histone acetyltransferase p300, may upregulate tumor suppressor pathways or reprogram oncogenic transcriptional networks.
Further, variability in therapeutic responses based on HER2 expression levels and mutation types highlights the influence of epigenetic regulation on treatment efficacy. CRISPRi and CRISPRa systems could be utilized to modulate chromatin accessibility and HER2 gene expression without introducing double-strand breaks, offering complementary strategies to pharmacologic HER2-targeted therapies. Overall, these findings establish a genomic and epigenetic framework for integrating CRISPR-based functional genomics into precision treatment strategies for HER2-mutant breast cancers.[27]
A randomized trial conducted by Roisin M et al. from March 2014 to October 2018 enrolled 608 patients with metastatic breast cancer to compare the efficacy of entinostat plus exemestane (EE) versus exemestane plus placebo (EP). A total of 305 patients were assigned to EE and 303 to EP, with a median age of 63 years (range 29 to 91). Most participants (60%) had visceral disease, and 84% were resistant to aromatase inhibitors (AI) in the metastatic setting. Prior therapies included chemotherapy (60%), CDK4/6 inhibitors (35%), and fulvestrant (30%), with baseline characteristics balanced across groups.
Regarding safety, patients in both arms received a median of three treatment cycles. Dose modifications were more frequent in the EE group (70% vs. 47%), and dose reductions occurred in 30% of EE patients compared to 3% in the EP group. Treatment discontinuation due to adverse events was higher in the EE arm (16% vs. 8%; P = 0.002). Grade 3–4 toxicities, particularly neutropenia (20%), hypophosphatemia (14%), and anemia (8%), were more common with EE, leading to an overall ≥ grade 3 toxicity rate of 51% compared to 16% in EP. There were 28 fatal adverse events, with four potentially related to treatment.
Efficacy outcomes revealed no significant advantage of EE over EP. Median PFS was 3.3 months for EE versus 3.1 months for EP (hazard ratio (HR) 0.87; 95% CI: 0.67–1.13; p = 0.30). Median OS was 23.4 months for EE versus 21.7 months for EP (HR 0.99; 95% CI: 0.82–1.21; p = 0.94), with similar objective response rates (5.8% EE vs. 5.6% EP). Despite a pharmacodynamic confirmation of entinostat’s activity—evidenced by a significant increase in lysine acetylation in peripheral blood mononuclear cells (median fold change ~1.5 vs. 1.0; p < 0.001), clinical outcomes did not improve.
The biological activity of entinostat emphasizes the potential of CRISPR-based epigenetic tools, such as CRISPR-dCas9 fused to histone deacetylases (HDACs) or histone acetyltransferases (HATs), to modulate specific chromatin loci more precisely than systemic HDAC inhibitors. For example, dCas9-HDAC3 or dCas9-p300 constructs could selectively repress ER-associated resistance genes or activate pro-apoptotic genes, potentially reducing systemic toxicity. Additionally, CRISPR screens could help identify predictive biomarkers of HDAC inhibition response, facilitating patient stratification and combinatorial strategies to overcome endocrine resistance.[28]
From December 2019 to November 2022, a study investigated by Abida W et al. in combination of ORIC-101, a selective glucocorticoid receptor (GR) antagonist, with enzalutamide in metastatic castration-resistant prostate cancer (mCRPC). Forty-one patients were enrolled, with dose escalation (n=10) and dose expansion (n=31) cohorts. Three dose levels of ORIC-101 (80 mg, 160 mg, 240 mg daily) combined with enzalutamide 160 mg daily were tested, establishing 240 mg as the recommended phase 2 dose (RP2D). No dose-limiting toxicities were observed, and pharmacodynamic assessments demonstrated consistent GR target gene suppression in peripheral blood mononuclear cells (PBMCs).
In the dose expansion cohort, 80.6% of patients experienced treatment-related adverse events, predominantly fatigue (38.7%) and nausea (29%). Interim analysis (n=28) showed a disease control rate (DCR) of 25% (80% CI: 10.7–44.9%), a PSA response rate of 0%, and an objective response rate (ORR) of 3.2%, with a median PFS of 4.1 months.
Comprehensive genomic profiling revealed that 39% of patients harbored AR resistance mutations or tumor suppressor gene losses, including tumour protein 53 (TP53), retinoblastoma 1 (RB1), and phosphatase and tensin homolog (PTEN), alongside MYC pathway activation. Patients lacking these alterations showed higher DCRs (42.1%) and better outcomes. This stratification underscores the utility of CRISPR-based approaches to edit or suppress resistance-driving mutations. Furthermore, emerging AR-dependent and AR-independent resistance pathways, such as PI3K-AKT-mTOR and Wnt signaling, represent CRISPR-editable targets in preclinical models for mCRPC.[29]
In a phase 1b/2a multicenter study of Horwitz SM et al., the safety and efficacy of duvelisib combined with romidepsin or bortezomib were evaluated in 105 patients with relapsed/refractory peripheral T-cell lymphoma (PTCL) and cutaneous T-cell lymphoma (CTCL). Arm A (n=66) received duvelisib plus romidepsin, and Arm B (n=32) received duvelisib plus bortezomib. In Arm A, the maximum tolerated dose was duvelisib 75 mg twice daily with romidepsin 10 mg/m². Arm B’s MTD was lower due to hepatotoxicity.
Duvelisib plus romidepsin yielded an ORR of 54% compared to 30% with the bortezomib combination. Specifically, in PTCL patients, the ORR was 56% with a CR rate of 44% (vs. 17% in Arm B, P = 0.0257). Molecular analyses identified somatic mutations in TET2, RHOA, and VAV1 enriched among responders. These mutations are attractive targets for CRISPR-mediated genome editing to dissect lymphomagenesis mechanisms. Resistance was associated with JAK1/JAK3 mutations and STAT pathway activation, corroborated by single-cell transcriptomics and CRISPR-based knockout studies demonstrating that TSC2 loss confers resistance via Akt/mTOR reactivation.[30]
A multicenter trial from July 2016 to March 2023 evaluated nemvaleukin alfa, an engineered IL-2 variant, alone or combined with pembrolizumab in advanced solid tumors. Among 243 patients, nemvaleukin monotherapy achieved an ORR of 10% (95% CI: 4 to 20%), with durable responses particularly in melanoma and RCC. Combination therapy resulted in an ORR of 13% (95% CI: 8 to 20%) and a median response duration of 65 weeks (95% CI: 21 to 160%). Immunomodulatory effects included expansion of CD8+ T cells and NK cells with minimal Treg induction.
This immune-activating profile positions nemvaleukin for potential enhancement via CRISPR-mediated engineering of T cells or NK cells, including knockout of inhibitory receptors like PD-1 or CTLA-4. CRISPR-based functional genomics could also uncover biomarkers predictive of response or resistance, guiding personalized immunotherapy approaches in IL-2 pathway modulation.[31]
| Study | Cancer type | Study phase | Intervention | Epigenetic or CRISPR relevance | CRISPR-based implications |
| Okines et al.[27] | HER2-mut MBC | Phase 2 | Tucatinib + Trastuzumab | HER2 mutations, chromatin dynamics | CRISPR-dCas9 for HER2 expression modulation and resistance mutation editing |
| Roisin et al.[28] | Metastatic breast cancer | RCT | Entinostat + Exemestane | HDAC inhibition, acetylation markers | CRISPR-HDAC/HAT fusions to modulate ER-resistance pathways |
| Abida et al.[29] | mCRPC | Phase 1/2 | ORIC-101 + Enzalutamide | GR pathway suppression | CRISPR editing of resistance mutations (TP53) |
| Horwitz et al.[30] | PTCL/CTCL | Phase 1b/2a | Duvelisib + Romidepsin/Bortezomib | Epigenetic mutations (TET2, RHOA) | Targeted knockout of JAK/STAT resistance nodes |
| Vaishampayan et al.[31] | Advanced solid tumors | Phase 1/2 | Nemvaleukin ± Pembrolizumab | Immunomodulation (IL-2 axis) | CRISPR enhancement of immune cells (PD-1 knockout) |
Table 1: Summary of included clinical trials and CRISPR-based implications
Discussion
CRISPR-based genome editing and epigenetic modulation have the potential to revolutionize cancer therapy by providing precise, programmable control over gene expression without introducing permanent changes to the DNA sequence. Traditional pharmacologic interventions targeting epigenetic mechanisms, such as histone deacetylase inhibitors, have demonstrated the potential to modify chromatin states and reverse gene silencing. However, their clinical application has been hindered by systemic toxicity, off-target effects, and limited therapeutic efficacy.
CRISPR technologies, particularly those involving catalytically dead Cas9 (dCas9) fused with epigenetic modifiers like HATs, HDACs, DNA methyltransferases, or demethylases, provide a method for site-specific modulation of gene expression. This approach allows for the reactivation of silenced tumor suppressor genes or suppression of oncogenes without inducing double-stranded DNA breaks, significantly lowering the risk of genomic instability. Studies have shown that dCas9-p300 fusion proteins can enhance transcriptional activation by locally acetylating histones, whereas dCas9-HDAC fusions can repress gene expression by promoting histone deacetylation. These capabilities open new avenues for reversing epigenetic silencing in cancer cells, particularly in cases where tumor suppressors are epigenetically inactivated.
Genomic profiling in various clinical trials reveals mutations and resistance pathways that are prime targets for CRISPR intervention. In HER2-mutant metastatic breast cancer, co-occurring mutations in CDH1 and PIK3CA influence treatment responses, suggesting potential for CRISPR-mediated correction of these resistance-associated mutations. In PTCL, mutations in TET2, RHOA, and VAV1 were enriched among responders to duvelisib and romidepsin combination therapy, indicating these as critical nodes for targeted genome editing to understand and potentially overcome resistance mechanisms.
CRISPR technologies also extend beyond DNA editing to RNA modulation through CRISPR-Cas13 systems, which can target and degrade specific RNA transcripts, offering a reversible and less permanent form of genetic intervention. However, significant barriers remain in translating CRISPR epigenetic editing to clinical settings, including the development of efficient and safe delivery systems capable of targeting tumor cells in vivo without eliciting off-target effects or immune responses.
Limitations
This literature review provides a comprehensive overview of recent clinical and translational studies on CRISPR-based epigenetic editing in cancer therapy. However, it has several limitations. The review excluded preclinical studies, in vitro research, and animal models, limiting the scope of mechanistic insights. This may result in underrepresentation of novel or emerging CRISPR modalities, particularly those involving newer Cas variants. Clinical trials were primarily pharmacologically driven, resulting in a hypothetical discussion about CRISPR’s application. The review also relied on a small number of trials, limiting the generalizability of findings across diverse cancer types. The heterogeneity of cancer mutations, epigenetic landscapes, and inter-individual variability in response to therapies also poses a challenge in drawing uniform conclusions on CRISPR’s therapeutic potential. Ethical concerns, regulatory barriers, and the absence of long-term safety data for CRISPR-based interventions in humans remain critical limitations for clinical translation.
Future studies
Future research should prioritize direct clinical trials that incorporate CRISPR-based epigenetic editing, particularly dCas9-fusion systems targeting histone modifications, DNA methylation, or RNA regulation in specific cancer subtypes. Expanding clinical research beyond breast, prostate, and T-cell lymphomas to include malignancies with well-characterized epigenetic dysregulation, such as glioblastoma or pancreatic cancer, would enhance the therapeutic breadth of CRISPR interventions.
Moreover, preclinical studies should focus on refining delivery platforms, such as tumor-specific lipid nanoparticles or exosome-mediated delivery, to improve the precision and safety of CRISPR components in vivo. Integrating AI-driven guide RNA design tools and high-throughput CRISPR screens can aid in minimizing off-target effects and optimizing target selection based on tumor-specific epigenetic vulnerabilities.
Another key direction involves combinatorial therapies, where CRISPR-mediated epigenetic editing is paired with immunotherapies, targeted agents, or chemotherapy to overcome resistance mechanisms and improve efficacy. Longitudinal studies to evaluate the durability, safety, and immunogenicity of CRISPR-based epigenetic interventions will also be essential for regulatory approval and clinical adoption.
Conclusion
CRISPR technologies have the potential to revolutionize cancer therapy by enabling precise and reversible modulation of gene expression through targeted epigenetic editing. The reviewed clinical studies, while pharmacologically driven, identify critical genetic and epigenetic landscapes where CRISPR interventions can be most impactful. Techniques utilizing dCas9-fusion proteins, CRISPRa/i, and RNA-targeting systems hold promise for reactivating tumor suppressor genes, silencing oncogenes, and circumventing drug resistance with minimal genomic disruption. Although significant hurdles remain, especially regarding delivery, specificity, and ethical concerns, ongoing advancements in guide RNA optimization, delivery systems, and computational biology are steadily paving the way for CRISPR’s clinical translation. Integrating CRISPR into precision oncology frameworks could ultimately lead to more effective, personalized, and less toxic cancer treatments.
References
- Pradeep KRK, Raj KB, Shrivastava SP, et al. An Observational Study on Drug-Related Problems in the Treatment of Cancer Patients. medtigo J Med. 2025;3(2):e3062326. doi:10.63096/medtigo3062326
Crossref - Irfan M, Majeed H, Iftikhar T, Ravi PK. A review on molecular scissoring with CRISPR/Cas9 genome editing technology. Toxicol Res (Camb). 2024;13(4):tfae105. doi:10.1093/toxres/tfae105
PubMed | Crossref | Google Scholar - Zhang C, Quan R, Wang J. Development and application of CRISPR/Cas9 technologies in genomic editing. Hum Mol Genet. 2018;27(R2):R79-88. doi:10.1093/hmg/ddy120
PubMed | Crossref | Google Scholar - Sun L, Zheng P, Sun J, Wendisch VF, Wang Y. Genome-scale CRISPRi screening: A powerful tool in engineering microbiology. Eng Microbiol. 2023;3(3):100089. doi:10.1016/j.engmic.2023.100089
PubMed | Crossref | Google Scholar - Nelles DA, Fang MY, O’Connell MR, et al. Programmable RNA Tracking in Live Cells with CRISPR/Cas9. Cell. 2016;165(2):488-496. doi:10.1016/j.cell.2016.02.054
PubMed | Crossref | Google Scholar - Zhen S, Lu JJ, Wang LJ, et al. In Vitro and In Vivo Synergistic Therapeutic Effect of Cisplatin with Human Papillomavirus16 E6/E7 CRISPR/Cas9 on Cervical Cancer Cell Line. Transl Oncol. 2016;9(6):498-504. doi:10.1016/j.tranon.2016.10.002
PubMed | Crossref | Google Scholar - Seeger C, Sohn JA. Complete Spectrum of CRISPR/Cas9-induced Mutations on HBV cccDNA. Mol Ther. 2016;24(7):1258-1266. doi:10.1038/mt.2016.94
PubMed | Crossref | Google Scholar - Kennedy EM, Bassit LC, Mueller H, et al. Suppression of hepatitis B virus DNA accumulation in chronically infected cells using a bacterial CRISPR/Cas RNA-guided DNA endonuclease. Virology. 2015;476:196-205. doi:10.1016/j.virol.2014.12.001
PubMed | Crossref | Google Scholar - Zhu W, Xie K, Xu Y, et al. CRISPR/Cas9 produces anti-hepatitis B virus effect in hepatoma cells and transgenic mouse. Virus Res. 2016;217:125-132. doi:10.1016/j.virusres.2016.04.003
PubMed | Crossref | Google Scholar - Yang L, Achreja A, Yeung TL, et al. Targeting Stromal Glutamine Synthetase in Tumors Disrupts Tumor Microenvironment-Regulated Cancer Cell Growth. Cell Metab. 2016;24(5):685-700. doi:10.1016/j.cmet.2016.10.011
PubMed | Crossref | Google Scholar - Zambirinis CP, Miller G. Cancer Manipulation of Host Physiology: Lessons from Pancreatic Cancer. Trends Mol Med. 2017;23(5):465-481. doi:10.1016/j.molmed.2017.03.003
PubMed | Crossref | Google Scholar - Zheng M, Li H, Gao Y, Brigstock DR, Gao R. Vitamin D3 analogue calcipotriol inhibits the profibrotic effects of transforming growth factor- β1 on pancreatic stellate cells. Eur J Pharmacol. 2023;957:176000. doi:10.1016/j.ejphar.2023.176000
PubMed | Crossref | Google Scholar - Neggers JE, Vercruysse T, Jacquemyn M, et al. Identifying drug-target selectivity of small-molecule CRM1/XPO1 inhibitors by CRISPR/Cas9 genome editing. Chem Biol. 2015;22(1):107-116. doi:10.1016/j.chembiol.2014.11.015
PubMed | Crossref | Google Scholar - Tzelepis K, Koike-Yusa H, De Braekeleer E, et al. A CRISPR Dropout Screen Identifies Genetic Vulnerabilities and Therapeutic Targets in Acute Myeloid Leukemia. Cell Rep. 2016;17(4):1193-1205. doi:10.1016/j.celrep.2016.09.079
PubMed | Crossref | Google Scholar - Topalian SL, Drake CG, Pardoll DM. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell. 2015;27(4):450-461. doi:10.1016/j.ccell.2015.03.001
PubMed | Crossref | Google Scholar - Ahmed S, Wan Azizan WAS, Nazri FR, Hatta MAM. Harnessing CRISPR/Cas9 system to engineer disease resistance in solanaceous crops: current progress and future prospects. Hortic Sci (Prague). 2025;52(1):1-14. doi:10.17221/19/2024-HORTSCI
Crossref | Google Scholar - Aubrey BJ, Kelly GL, Kueh AJ, et al. An inducible lentiviral guide RNA platform enables the identification of tumor-essential genes and tumor-promoting mutations in vivo. Cell Rep. 2015;10(8):1422-1432. doi:10.1016/j.celrep.2015.02.002
PubMed | Crossref | Google Scholar - Antal CE, Hudson AM, Kang E, et al. Cancer-associated protein kinase C mutations reveal kinase’s role as tumor suppressor. Cell. 2015;160(3):489-502. doi:10.1016/j.cell.2015.01.001
PubMed | Crossref | Google Scholar - Hoppe-Seyler K, Bossler F, Braun JA, Herrmann AL, Hoppe-Seyler F. The HPV E6/E7 Oncogenes: Key Factors for Viral Carcinogenesis and Therapeutic Targets. Trends Microbiol. 2018;26(2):158-168. doi:10.1016/j.tim.2017.07.007
PubMed | Crossref | Google Scholar - Anand U, Dey A, Chandel AKS, et al. Cancer chemotherapy and beyond: Current status, drug candidates, associated risks and progress in targeted therapeutics. Genes Dis. 2022;10(4):1367-1401. doi:10.1016/j.gendis.2022.02.007
PubMed | Crossref | Google Scholar - Roychowdhury S, Chinnaiyan AM. Translating cancer genomes and transcriptomes for precision oncology. CA Cancer J Clin. 2016;66(1):75-88. doi:10.3322/caac.21329
PubMed | Crossref | Google Scholar - Maresso KC, Tsai KY, Brown PH, Szabo E, Lippman S, Hawk ET. Molecular cancer prevention: Current status and future directions. CA Cancer J Clin. 2015;65(5):345-383. doi:10.3322/caac.21287
PubMed | Crossref | Google Scholar - Sachdeva M, Sachdeva N, Pal M, et al. CRISPR/Cas9: molecular tool for gene therapy to target genome and epigenome in the treatment of lung cancer. Cancer Gene Ther. 2015;22(11):509-517. doi:10.1038/cgt.2015.54
PubMed | Crossref | Google Scholar - Ansori AN, Antonius Y, Susilo RJ, et al. Application of CRISPR-Cas9 genome editing technology in various fields: A review. Narra J. 2023;3(2):e184. doi:10.52225/narra.v3i2.184
PubMed | Crossref | Google Scholar - Lee M. Deep learning in CRISPR-Cas systems: a review of recent studies. Front Bioeng Biotechnol. 2023;11:1226182. doi:10.3389/fbioe.2023.1226182
PubMed | Crossref | Google Scholar - Charlier J, Nadon R, Makarenkov V. Accurate deep learning off-target prediction with novel sgRNA-DNA sequence encoding in CRISPR-Cas9 gene editing. Bioinformatics. 2021;37(16):2299-2307. doi:10.1093/bioinformatics/btab112
PubMed | Crossref | Google Scholar - Okines AFC, Curigliano G, Mizuno N, et al. Tucatinib and trastuzumab in HER2-mutated metastatic breast cancer: a phase 2 basket trial. Nat Med. 2025;31(3):909-916. doi:10.1038/s41591-024-03462-0
PubMed | Crossref | Google Scholar - Connolly RM, Zhao F, Miller KD, et al. E2112: Randomized Phase III Trial of Endocrine Therapy Plus Entinostat or Placebo in Hormone Receptor-Positive Advanced Breast Cancer. A Trial of the ECOG-ACRIN Cancer Research Group. J Clin Oncol. 2021;39(28):3171-3181. doi:10.1200/JCO.21.00944
PubMed | Crossref | Google Scholar - Abida W, Hahn AW, Shore N, et al. Phase I Study of ORIC-101, a Glucocorticoid Receptor Antagonist, in Combination with Enzalutamide in Patients with Metastatic Castration-resistant Prostate Cancer Progressing on Enzalutamide. Clin Cancer Res. 2024;30(6):1111-1120. doi:10.1158/1078-0432.CCR-23-3508
PubMed | Crossref | Google Scholar - Horwitz SM, Nirmal AJ, Rahman J, et al. Duvelisib plus romidepsin in relapsed/refractory T cell lymphomas: a phase 1b/2a trial. Nat Med. 2024;30(9):2517-2527. doi:10.1038/s41591-024-03076-6
PubMed | Crossref | Google Scholar - Vaishampayan UN, Muzaffar J, Winer I, et al. Nemvaleukin alfa, a modified interleukin-2 cytokine, as monotherapy and with pembrolizumab in patients with advanced solid tumors (ARTISTRY-1). J Immunother Cancer. 2024;12(11):e010143. doi:10.1136/jitc-2024-010143
PubMed | Crossref | Google Scholar
Acknowledgments
Not reported
Funding
No funding
Author Information
Corresponding Author:
Samatha Ampeti, PhD
Independent Researcher, Department of Content
medtigo India Pvt Ltd, Pune, India
Email: ampetisamatha9@gmail.com
Co-Authors:
Patel Nirali Kirankumar, Mansi Srivastava, Raziya Begum Sheikh, Shubham Ravindra Sali, Sonam Shashikala B V
Independent Researcher, Department of Content
medtigo India Pvt Ltd, Pune, India
Authors Contributions
All authors contributed to the conceptualization, investigation, and data curation by acquiring and critically reviewing the selected articles. They were collectively involved in the writing – original draft preparation and writing – review & editing to refine the manuscript. Additionally, all authors participated in the supervision of the work, ensuring accuracy and completeness. The final manuscript was approved by all named authors for submission to the journal.
Ethical Approval
Not applicable
Conflict of Interest Statement
None
Guarantor
None
DOI
Cite this Article
Patel NK, Samatha A, Mansi S, Raziya BS, Shubham RS, Sonam SBV. Targeted Epigenome Editing with CRISPR Technologies: Transforming Cancer Treatment Strategies. medtigo J Pharmacol. 2025;2(3):e3061239. doi:10.63096/medtigo3061239 Crossref









